Abstract
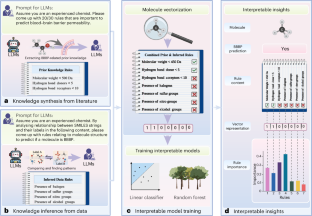
Large language models (LLMs) are a form of artificial intelligence system encapsulating vast knowledge in the form of natural language. These systems are adept at numerous complex tasks including creative writing, storytelling, translation, question-answering, summarization and computer code generation. Although LLMs have seen initial applications in natural sciences, their potential for driving scientific discovery remains largely unexplored. In this work, we introduce LLM4SD, a framework designed to harness LLMs for driving scientific discovery in molecular property prediction by synthesizing knowledge from literature and inferring knowledge from scientific data. LLMs synthesize knowledge by extracting established information from scientific literature, such as molecular weight being key to predicting solubility. For inference, LLMs identify patterns in molecular data, particularly in Simplified Molecular Input Line Entry System-encoded structures, such as halogen-containing molecules being more likely to cross the blood–brain barrier. This information is presented as interpretable knowledge, enabling the transformation of molecules into feature vectors. By using these features with interpretable models such as random forest, LLM4SD can outperform the current state of the art across a range of benchmark tasks for predicting molecular properties. We foresee it providing interpretable and potentially new insights, aiding scientific discovery in molecular property prediction.
This is a preview of subscription content, access via your institution
Access options
/* style specs end */
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
/* style specs start */
style {
display: none !important;
}
.LiveAreaSection * {
align-content: stretch;
align-items: stretch;
align-self: auto;
animation-delay: 0s;
animation-direction: normal;
animation-duration: 0s;
animation-fill-mode: none;
animation-iteration-count: 1;
animation-name: none;
animation-play-state: running;
animation-timing-function: ease;
azimuth: center;
backface-visibility: visible;
background-attachment: scroll;
background-blend-mode: normal;
background-clip: borderBox;
background-color: transparent;
background-image: none;
background-origin: paddingBox;
background-position: 0 0;
background-repeat: repeat;
background-size: auto auto;
block-size: auto;
border-block-end-color: currentcolor;
border-block-end-style: none;
border-block-end-width: medium;
border-block-start-color: currentcolor;
border-block-start-style: none;
border-block-start-width: medium;
border-bottom-color: currentcolor;
border-bottom-left-radius: 0;
border-bottom-right-radius: 0;
border-bottom-style: none;
border-bottom-width: medium;
border-collapse: separate;
border-image-outset: 0s;
border-image-repeat: stretch;
border-image-slice: 100%;
border-image-source: none;
border-image-width: 1;
border-inline-end-color: currentcolor;
border-inline-end-style: none;
border-inline-end-width: medium;
border-inline-start-color: currentcolor;
border-inline-start-style: none;
border-inline-start-width: medium;
border-left-color: currentcolor;
border-left-style: none;
border-left-width: medium;
border-right-color: currentcolor;
border-right-style: none;
border-right-width: medium;
border-spacing: 0;
border-top-color: currentcolor;
border-top-left-radius: 0;
border-top-right-radius: 0;
border-top-style: none;
border-top-width: medium;
bottom: auto;
box-decoration-break: slice;
box-shadow: none;
box-sizing: border-box;
break-after: auto;
break-before: auto;
break-inside: auto;
caption-side: top;
caret-color: auto;
clear: none;
clip: auto;
clip-path: none;
color: initial;
column-count: auto;
column-fill: balance;
column-gap: normal;
column-rule-color: currentcolor;
column-rule-style: none;
column-rule-width: medium;
column-span: none;
column-width: auto;
content: normal;
counter-increment: none;
counter-reset: none;
cursor: auto;
display: inline;
empty-cells: show;
filter: none;
flex-basis: auto;
flex-direction: row;
flex-grow: 0;
flex-shrink: 1;
flex-wrap: nowrap;
float: none;
font-family: initial;
font-feature-settings: normal;
font-kerning: auto;
font-language-override: normal;
font-size: medium;
font-size-adjust: none;
font-stretch: normal;
font-style: normal;
font-synthesis: weight style;
font-variant: normal;
font-variant-alternates: normal;
font-variant-caps: normal;
font-variant-east-asian: normal;
font-variant-ligatures: normal;
font-variant-numeric: normal;
font-variant-position: normal;
font-weight: 400;
grid-auto-columns: auto;
grid-auto-flow: row;
grid-auto-rows: auto;
grid-column-end: auto;
grid-column-gap: 0;
grid-column-start: auto;
grid-row-end: auto;
grid-row-gap: 0;
grid-row-start: auto;
grid-template-areas: none;
grid-template-columns: none;
grid-template-rows: none;
height: auto;
hyphens: manual;
image-orientation: 0deg;
image-rendering: auto;
image-resolution: 1dppx;
ime-mode: auto;
inline-size: auto;
isolation: auto;
justify-content: flexStart;
left: auto;
letter-spacing: normal;
line-break: auto;
line-height: normal;
list-style-image: none;
list-style-position: outside;
list-style-type: disc;
margin-block-end: 0;
margin-block-start: 0;
margin-bottom: 0;
margin-inline-end: 0;
margin-inline-start: 0;
margin-left: 0;
margin-right: 0;
margin-top: 0;
mask-clip: borderBox;
mask-composite: add;
mask-image: none;
mask-mode: matchSource;
mask-origin: borderBox;
mask-position: 0 0;
mask-repeat: repeat;
mask-size: auto;
mask-type: luminance;
max-height: none;
max-width: none;
min-block-size: 0;
min-height: 0;
min-inline-size: 0;
min-width: 0;
mix-blend-mode: normal;
object-fit: fill;
object-position: 50% 50%;
offset-block-end: auto;
offset-block-start: auto;
offset-inline-end: auto;
offset-inline-start: auto;
opacity: 1;
order: 0;
orphans: 2;
outline-color: initial;
outline-offset: 0;
outline-style: none;
outline-width: medium;
overflow: visible;
overflow-wrap: normal;
overflow-x: visible;
overflow-y: visible;
padding-block-end: 0;
padding-block-start: 0;
padding-bottom: 0;
padding-inline-end: 0;
padding-inline-start: 0;
padding-left: 0;
padding-right: 0;
padding-top: 0;
page-break-after: auto;
page-break-before: auto;
page-break-inside: auto;
perspective: none;
perspective-origin: 50% 50%;
pointer-events: auto;
position: static;
quotes: initial;
resize: none;
right: auto;
ruby-align: spaceAround;
ruby-merge: separate;
ruby-position: over;
scroll-behavior: auto;
scroll-snap-coordinate: none;
scroll-snap-destination: 0 0;
scroll-snap-points-x: none;
scroll-snap-points-y: none;
scroll-snap-type: none;
shape-image-threshold: 0;
shape-margin: 0;
shape-outside: none;
tab-size: 8;
table-layout: auto;
text-align: initial;
text-align-last: auto;
text-combine-upright: none;
text-decoration-color: currentcolor;
text-decoration-line: none;
text-decoration-style: solid;
text-emphasis-color: currentcolor;
text-emphasis-position: over right;
text-emphasis-style: none;
text-indent: 0;
text-justify: auto;
text-orientation: mixed;
text-overflow: clip;
text-rendering: auto;
text-shadow: none;
text-transform: none;
text-underline-position: auto;
top: auto;
touch-action: auto;
transform: none;
transform-box: borderBox;
transform-origin: 50% 50%0;
transform-style: flat;
transition-delay: 0s;
transition-duration: 0s;
transition-property: all;
transition-timing-function: ease;
vertical-align: baseline;
visibility: visible;
white-space: normal;
widows: 2;
width: auto;
will-change: auto;
word-break: normal;
word-spacing: normal;
word-wrap: normal;
writing-mode: horizontalTb;
z-index: auto;
-webkit-appearance: none;
-moz-appearance: none;
-ms-appearance: none;
appearance: none;
margin: 0;
}
.LiveAreaSection {
width: 100%;
}
.LiveAreaSection .login-option-buybox {
display: block;
width: 100%;
font-size: 17px;
line-height: 30px;
color: #222;
padding-top: 30px;
font-family: Harding, Palatino, serif;
}
.LiveAreaSection .additional-access-options {
display: block;
font-weight: 700;
font-size: 17px;
line-height: 30px;
color: #222;
font-family: Harding, Palatino, serif;
}
.LiveAreaSection .additional-login > li:not(:first-child)::before {
transform: translateY(-50%);
content: “”;
height: 1rem;
position: absolute;
top: 50%;
left: 0;
border-left: 2px solid #999;
}
.LiveAreaSection .additional-login > li:not(:first-child) {
padding-left: 10px;
}
.LiveAreaSection .additional-login > li {
display: inline-block;
position: relative;
vertical-align: middle;
padding-right: 10px;
}
.BuyBoxSection {
display: flex;
flex-wrap: wrap;
flex: 1;
flex-direction: row-reverse;
margin: -30px -15px 0;
}
.BuyBoxSection .box-inner {
width: 100%;
height: 100%;
padding: 30px 5px;
display: flex;
flex-direction: column;
justify-content: space-between;
}
.BuyBoxSection p {
margin: 0;
}
.BuyBoxSection .readcube-buybox {
background-color: #f3f3f3;
flex-shrink: 1;
flex-grow: 1;
flex-basis: 255px;
background-clip: content-box;
padding: 0 15px;
margin-top: 30px;
}
.BuyBoxSection .subscribe-buybox {
background-color: #f3f3f3;
flex-shrink: 1;
flex-grow: 4;
flex-basis: 300px;
background-clip: content-box;
padding: 0 15px;
margin-top: 30px;
}
.BuyBoxSection .subscribe-buybox-nature-plus {
background-color: #f3f3f3;
flex-shrink: 1;
flex-grow: 4;
flex-basis: 100%;
background-clip: content-box;
padding: 0 15px;
margin-top: 30px;
}
.BuyBoxSection .title-readcube,
.BuyBoxSection .title-buybox {
display: block;
margin: 0;
margin-right: 10%;
margin-left: 10%;
font-size: 24px;
line-height: 32px;
color: #222;
text-align: center;
font-family: Harding, Palatino, serif;
}
.BuyBoxSection .title-asia-buybox {
display: block;
margin: 0;
margin-right: 5%;
margin-left: 5%;
font-size: 24px;
line-height: 32px;
color: #222;
text-align: center;
font-family: Harding, Palatino, serif;
}
.BuyBoxSection .asia-link,
.Link-328123652,
.Link-2926870917,
.Link-2291679238,
.Link-595459207 {
color: #069;
cursor: pointer;
text-decoration: none;
font-size: 1.05em;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 1.05em6;
}
.BuyBoxSection .access-readcube {
display: block;
margin: 0;
margin-right: 10%;
margin-left: 10%;
font-size: 14px;
color: #222;
padding-top: 10px;
text-align: center;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 20px;
}
.BuyBoxSection ul {
margin: 0;
}
.BuyBoxSection .link-usp {
display: list-item;
margin: 0;
margin-left: 20px;
padding-top: 6px;
list-style-position: inside;
}
.BuyBoxSection .link-usp span {
font-size: 14px;
color: #222;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 20px;
}
.BuyBoxSection .access-asia-buybox {
display: block;
margin: 0;
margin-right: 5%;
margin-left: 5%;
font-size: 14px;
color: #222;
padding-top: 10px;
text-align: center;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 20px;
}
.BuyBoxSection .access-buybox {
display: block;
margin: 0;
margin-right: 10%;
margin-left: 10%;
font-size: 14px;
color: #222;
opacity: 0.8px;
padding-top: 10px;
text-align: center;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 20px;
}
.BuyBoxSection .price-buybox {
display: block;
font-size: 30px;
color: #222;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
padding-top: 30px;
text-align: center;
}
.BuyBoxSection .price-buybox-to {
display: block;
font-size: 30px;
color: #222;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
text-align: center;
}
.BuyBoxSection .price-info-text {
font-size: 16px;
padding-right: 10px;
color: #222;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
}
.BuyBoxSection .price-value {
font-size: 30px;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
}
.BuyBoxSection .price-per-period {
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
}
.BuyBoxSection .price-from {
font-size: 14px;
padding-right: 10px;
color: #222;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 20px;
}
.BuyBoxSection .issue-buybox {
display: block;
font-size: 13px;
text-align: center;
color: #222;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 19px;
}
.BuyBoxSection .no-price-buybox {
display: block;
font-size: 13px;
line-height: 18px;
text-align: center;
padding-right: 10%;
padding-left: 10%;
padding-bottom: 20px;
padding-top: 30px;
color: #222;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
}
.BuyBoxSection .vat-buybox {
display: block;
margin-top: 5px;
margin-right: 20%;
margin-left: 20%;
font-size: 11px;
color: #222;
padding-top: 10px;
padding-bottom: 15px;
text-align: center;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: 17px;
}
.BuyBoxSection .tax-buybox {
display: block;
width: 100%;
color: #222;
padding: 20px 16px;
text-align: center;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
line-height: NaNpx;
}
.BuyBoxSection .button-container {
display: flex;
padding-right: 20px;
padding-left: 20px;
justify-content: center;
}
.BuyBoxSection .button-container > * {
flex: 1px;
}
.BuyBoxSection .button-container > a:hover,
.Button-505204839:hover,
.Button-1078489254:hover,
.Button-2737859108:hover {
text-decoration: none;
}
.BuyBoxSection .btn-secondary {
background: #fff;
}
.BuyBoxSection .button-asia {
background: #069;
border: 1px solid #069;
border-radius: 0;
cursor: pointer;
display: block;
padding: 9px;
outline: 0;
text-align: center;
text-decoration: none;
min-width: 80px;
margin-top: 75px;
}
.BuyBoxSection .button-label-asia,
.ButtonLabel-3869432492,
.ButtonLabel-3296148077,
.ButtonLabel-1636778223 {
display: block;
color: #fff;
font-size: 17px;
line-height: 20px;
font-family: -apple-system, BlinkMacSystemFont, “Segoe UI”, Roboto,
Oxygen-Sans, Ubuntu, Cantarell, “Helvetica Neue”, sans-serif;
text-align: center;
text-decoration: none;
cursor: pointer;
}
.Button-505204839,
.Button-1078489254,
.Button-2737859108 {
background: #069;
border: 1px solid #069;
border-radius: 0;
cursor: pointer;
display: block;
padding: 9px;
outline: 0;
text-align: center;
text-decoration: none;
min-width: 80px;
max-width: 320px;
margin-top: 20px;
}
.Button-505204839 .btn-secondary-label,
.Button-1078489254 .btn-secondary-label,
.Button-2737859108 .btn-secondary-label {
color: #069;
}
.uList-2102244549 {
list-style: none;
padding: 0;
margin: 0;
}
/* style specs end */

Similar content being viewed by others
Leveraging large language models for predictive chemistry
Augmenting large language models with chemistry tools

Large language models surpass human experts in predicting neuroscience results
Data availability
The split datasets utilized in this study are entirely open source and have been made publicly available to ensure straightforward replication of our findings. We have provided presplit datasets for a variety of tasks, including BBBP, ClinTox, Tox21 (12 subtasks), SIDER (27 subtasks), HIV, BACE, ESOL, Lipophilicity, FreeSolv and QM9 (12 subtasks). These datasets are divided into training, validation and test sets based on our experimental settings. You can access them at the following GitHub repository: https://github.com/zyzisastudyreallyhardguy/LLM4SD/tree/main/scaffold_datasets. For the original raw datasets provided by MoleculeNet9, the corresponding links are as follows: BBBP, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/BBBP.csv; ClinTox, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/clintox.csv.gz; Tox21, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/tox21.csv.gz; SIDER, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/sider.csv.gz; HIV, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/HIV.csv; BACE, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/bace.csv; ESOL, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/delaney-processed.csv; FreeSolv, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/SAMPL.csv; Lipophilicity, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/Lipophilicity.csv; QM9, https://deepchemdata.s3-us-west-1.amazonaws.com/datasets/qm9.csv. Source data are provided with this paper.
Code availability
In our commitment to transparency and reproducibility, we have released our code showing our implementation. This encompasses methodologies for literature knowledge mining, knowledge inference rule mining and interpretable model training. Throughout this work, we have employed several open-source libraries, including Hugging Face, numpy, rdkit, pytorch, scipy, bitsandbytes and accelerate. The GitHub link of the model is https://github.com/zyzisastudyreallyhardguy/LLM4SD (https://doi.org/10.5281/zenodo.13986921)43. Furthermore, we are in the process of deploying a website to facilitate scientists in utilizing LLM4SD. The site features three core functionalities for scientific users: knowledge synthesis, knowledge inference and prediction with explanations. Examples of user interactions with the website can be found in the Supplementary Information.
References
-
Bloom, N., Jones, C. I., Reenen, J. & Webb, M. Are ideas getting harder to find? Am. Econ. Rev. 110, 1104–1144 (2020).
Google Scholar
-
Wang, H. et al. Scientific discovery in the age of artificial intelligence. Nature 620, 47–60 (2023).
Google Scholar
-
Frank, M. Baby steps in evaluating the capacities of large language models. Nat. Rev. Psychol. 2, 451–452 (2023).
-
Brown, T. et al. Language models are few-shot learners. Adv. Neural Inf. Process Syst. 33, 1877–901 (2020).
Google Scholar
-
Achiam, J. et al. Gpt-4 technical report. Preprint at https://arxiv.org/abs/2303.08774 (2023).
-
Jiang, L. Y. et al. Health system-scale language models are all-purpose prediction engines. Nature 619, 357–362 (2023).
-
Singhal, K. et al. Large language models encode clinical knowledge. Nature 620, 172–180 (2023).
-
Mirza, A. et al. Are large language models superhuman chemists? Preprint at https://arxiv.org/abs/2404.01475 (2024).
-
Wu, Z. et al. MoleculeNet: a benchmark for molecular machine learning. Chem. Sci. 9, 513–530 (2018).
Google Scholar
-
Taylor, R. et al. Galactica: a large language model for science. Preprint at https://arxiv.org/abs/2211.09085 (2022).
-
Almazrouei, E. et al. The Falcon series of open language models. Preprint at https://arxiv.org/abs/2311.16867 (2023).
-
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 64, 4–17 (2012).
Google Scholar
-
Landrum, G. et al. rdkit/rdkit: 2024_09_5 (Q3 2024) Release (Release_2024_09_5). Zenodo https://doi.org/10.5281/zenodo.14779836 (2025).
-
Hu, W. et al. Strategies for pre-training graph neural networks. In Proc. International Conference on Learning Representations (2020).
-
You, Y. et al. Graph contrastive learning with augmentations. Adv. Neural Inf. Process. Sys. 33, 5812–5823 (2020).
Google Scholar
-
Wang, Y., Wang, J., Cao, Z. & Farimani, A. B. Molecular contrastive learning of representations via graph neural networks. Nat. Mach. Intell. 4, 279–287 (2022).
Google Scholar
-
Stärk, H. et al. 3D Infomax improves GNNs for molecular property prediction. In Proc. International Conference on Machine Learning (eds Chaudhuri, K. et al.) 20479–20502 (PMLR, 2022).
-
Liu, S. et al. Pre-training molecular graph representation with 3D geometry. In Proc. 10th International Conference on Learning Representations (2022).
-
Xia, J. et al. Mole-bert: rethinking pre-training graph neural networks for molecules. In Proc. 11th International Conference on Learning Representations (2023).
-
Rong, Y. et al. Self-supervised graph transformer on large-scale molecular data. Adv. Neural Inf. Process. Sys. 33, 12559–12571 (2020).
Google Scholar
-
Zhou, G. et al. Uni-mol: a universal 3D molecular representation learning framework. In Proc. 11th International Conference on Learning Representations (2023).
-
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 50, 742–754 (2010).
Google Scholar
-
Stokes, J. M. et al. A deep learning approach to antibiotic discovery. Cell 180, 688–702.e13 (2020).
Google Scholar
-
Wong, F. et al. Discovery of a structural class of antibiotics with explainable deep learning. Nature 626, 177–185 (2024).
Google Scholar
-
Zhang, D. et al. Chemllm: a chemical large language model. Preprint at https://arxiv.org/abs/2402.06852 (2024).
-
Zhao, Z. et al. ChemDFM: a large language foundation model for chemistry. In 38th Conference on Neural Information Processing Systems, Foundation Models for Science: Progress, Opportunities, and Challenges (NeurIPS, 2024).
-
Cai, Z. et al. Internlm2 technical report. Preprint at https://arxiv.org/abs/2403.17297 (2024).
-
Touvron, H. et al. Llama: open and efficient foundation language models. Preprint at https://arxiv.org/abs/2302.13971 (2023).
-
Haque, M. & Li, S. Exploring ChatGPT and its impact on society. AI Ethics https://doi.org/10.1007/s43681-024-00435-4 (2024).
-
Wei, J. et al. Emergent abilities of large language models. Transact. Mach. Learn. Res. https://openreview.net/pdf?id=yzkSU5zdwD (2022).
-
McKnight, P. E. & Najab, J. in The Corsini Encyclopedia of Psychology (eds Weiner, I. B. & Craighead, W. E.) (Wiley, 2010).
-
Subramanian, G., Ramsundar, B., Pande, V. & Denny, R. A. Computational modeling of β-secretase 1 (BACE-1) inhibitors using ligand based approaches. J. Chem. Inf. Model. 56, 1936–1949 (2016).
Google Scholar
-
Wager, T. Defining desirable central nervous system drug space through the alignment of molecular properties, in vitro adme, and safety attributes. ACS Chem. Neurosci. 1, 420–434 (2010).
Google Scholar
-
Wager, T., Hou, X., Verhoest, P. & Villalobos, A. Moving beyond rules: the development of a central nervous system multiparameter optimization (cns mpo) approach to enable alignment of druglike properties. ACS Chem. Neurosci. 1, 435–449 (2010).
Google Scholar
-
Geldenhuys, W., Mohammad, A., Adkins, C. & Lockman, P. Molecular determinants of blood–brain barrier permeation. Ther. Deliv. 6, 961–971 (2015).
Google Scholar
-
Liu, N. F. et al. Lost in the middle: how language models use long contexts. Trans. Assoc. Comput. Linguist. 12, 157–173 (2024).
Google Scholar
-
Qin, G., Feng, Y. & Van Durme, B. The NLP task effectiveness of long-range transformers. In Proc. 17th Conference of the European Chapter of the Association for Computational Linguistics (eds Vlachos, A. & Augenstein, I.) 3774–3790 (ACL, 2023).
-
The UniProt Consortium. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
Google Scholar
-
Benson, D. A. et al. GenBank. Nucleic Acids Res. 41, D36–D42 (2013).
Google Scholar
-
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Google Scholar
-
Park, Y. J. et al. Can chatgpt be used to generate scientific hypotheses? J. Materiomics 10, 578–584 (2024).
Google Scholar
-
Honda, S., Shi, S. & Ueda, H. R. Smiles transformer: pre-trained molecular fingerprint for low data drug discovery. Preprint at https://arxiv.org/abs/1911.04738 (2019).
-
zyzisastudyreallyhardguy & Ju, J. Code repository LLM4SD: release v.1.0. Zenodo https://doi.org/10.5281/zenodo.13986921 (2024).
-
Student. The probable error of a mean. Biometrika 6, 1–25 (1908).
Acknowledgements
H.Y.K.’s scholarship is supported by the Australian Government Research Training Programme (RTP) Scholarship and Monash University as a cocontribution to Australian Research Council grant no. ARC DP210100072. L.T.M.’s, G.I.W.’s and A.T.N.N.’s research into AI applications for drug discovery is supported by a National Health and Medical Research Council (NHMRC) of Australia Ideas grant (grant no. APP2013629). L.T.M.’s research is also supported by the National Heart Foundation of Australia (grant no. 101857). L.T.M.’s and A.T.N.N.’s research is also funded by the NHMRC of Australia and the Department of Health and Aged Care through the Medical Research Future Fund (MRFF) Stem Cell Therapies Mission (grant no. MRF2015957). Computational resources were generously provided by the Nectar Research Cloud, a collaborative Australian research platform supported by the NCRIS-funded Australian Research Data Commons (ARDC) and the MASSIVE HPC facility. We also gratefully acknowledge the support of the Griffith University eResearch Service & Specialized Platforms Team and the use of the High-Performance Computing Cluster ‘Gowonda’. S.P. is supported by ARC Future Fellowship (grant no. FT210100097) and ARC grant no. DP240101547.
Author information
Authors and Affiliations
Contributions
S.P. and G.I.W. supervised the project. Y.Z., H.Y.K. and J.J. contributed to the conception and design of the work. Y.Z., H.Y.K. and J.J. contributed to the technical implementation. Y.Z., H.Y.K. and J.J. prepared the figures. Y.Z. contributed to the design of the web-based application. A.T.N.N. and L.T.M. provided domain expertise for the literature review and validation of rules. Y.Z., H.Y.K., A.T.N.N. and L.T.M. contributed to the design of the rule validation test. All authors edited and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
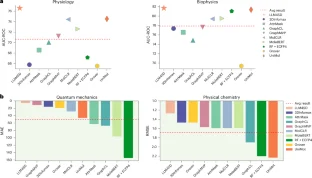
Extended Data Fig. 1 Detailed performance comparison between ‘LLM4SD’ and nine baselines in the physiology domain.
The red dashed line shows the average result across all methods. Each marker’s error bar denotes the method’s standard deviation, which is obtained via 10 runs. LLM4SD outperformed other models in 3 out of 4 datasets using the AUC-ROC metric and consistently surpassing the average across all datasets. The results for Tox21 and SIDER are average scores from 12 and 27 tasks respectively (see Extended Data Figs. 2 and 3 for detailed breakdown).
Source data
Extended Data Fig. 2 Detailed performance comparison between ‘LLM4SD’ and nine baselines on Tox21 Dataset.
The red dashed line shows the average result across all methods. Each marker’s error bar denotes the method’s standard deviation, which is obtained via 10 runs. LLM4SD ranks among the top three methods in 8 out of 12 tasks and consistently outperformed the average in all tasks.
Source data
Extended Data Fig. 3 Detailed performance comparison between ‘LLM4SD’ and nine baselines on Sider Dataset.
The red dashed line shows the average result across all methods. Each marker’s error bar denotes the method’s standard deviation, which is obtained via 10 runs. LLM4SD ranks among the top three methods in 22 out of 27 tasks, and consistently outperforms the average in all tasks with the exception of the ‘Psychiatric disorders’ task.
Source data
Extended Data Fig. 4 Detailed performance comparison between ‘LLM4SD’ and nine baselines in the biophysics domain.
The red dashed line shows the average result across all methods, in terms of AUC-ROC. Each marker’s error bar denotes the method’s standard deviation, which is obtained via 10 runs. LLM4SD outperformed the top-performing baseline by roughly 1% on the HIV dataset and closely matched the best performing method, UniMol. In both cases, LLM4SD delivered a visibly superior outcome compared to the average performance.
Source data
Extended Data Fig. 5 Detailed performance comparison between ‘LLM4SD’ and nine baselines in the physical chemistry domain.
The red dashed line shows the average result across all methods. The physical chemistry domain encompasses three datasets: the ESOL dataset with 1,128 instances, the FreeSolv dataset with 642 instances, and the Lipophilicity dataset comprising 4,200 compounds. Each marker’s error bar represents the method’s standard deviation, calculated based on 10 independent runs (n=10). These data points are overlaid on the plot in grey colour. LLM4SD substantially outperformed all baseline methods on ESOL, demonstrating a 57% improvement over the average outcome for that dataset, and achieved state-of-the-art results on the additional datasets, FreeSolv and Lipophilicity.
Source data
Extended Data Fig. 6 Detailed performance comparison between ‘LLM4SD’ and nine baselines in the quantum mechanics domain.
The red dashed line shows the average result across all methods. The quantum mechanics domain includes the QM9 datasets with 12 subtasks, comprising 133,885 instances. Each marker’s error bar denotes the method’s standard deviation, which is obtained via 10 runs (n=10). These data points are overlaid on the plot in grey colour. LLM4SD excelled in predicting properties such as U0, U, H, and G, showing substantial enhancements. In other tasks, the results from LLM4SD were comparable to the average of all methods.
Source data
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and Figs. 1–9.
Reporting Summary
Source data
Source Data Fig. 2
Source data for plotting the figure.
Source Data Fig. 3
Source data for plotting the figure.
Source Data Fig. 4
Source data for plotting the figure.
Source Data Extended Data Fig. 1
Source data for plotting the figure.
Source Data Extended Data Fig. 2
Source data for plotting the figure.
Source Data Extended Data Fig. 3
Source data for plotting the figure.
Source Data Extended Data Fig. 4
Source data for plotting the figure.
Source Data Extended Data Fig. 5
Source data for plotting the figure.
Source Data Extended Data Fig. 6
Source data for plotting the figure.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
Reprints and permissions
About this article
Cite this article
Zheng, Y., Koh, H.Y., Ju, J. et al. Large language models for scientific discovery in molecular property prediction.
Nat Mach Intell (2025). https://doi.org/10.1038/s42256-025-00994-z
-
Received: 24 October 2023
-
Accepted: 15 January 2025
-
Published: 25 February 2025
-
DOI: https://doi.org/10.1038/s42256-025-00994-z